github.com/StephLocke | itsalocke.com | T: SteffLocke
R is an integrated suite of software facilities for data manipulation, calculation and graphical display
I lied…
Please note these instructions were tested on a Windows 8.1 clean VM from Azure on 20/07/2015 - updates to r, rstudio, packages, or the training package could cause drift between what works and what doesn't and you'll have to wing it. If you do have to wing it, let me know and I'll update my instructions!
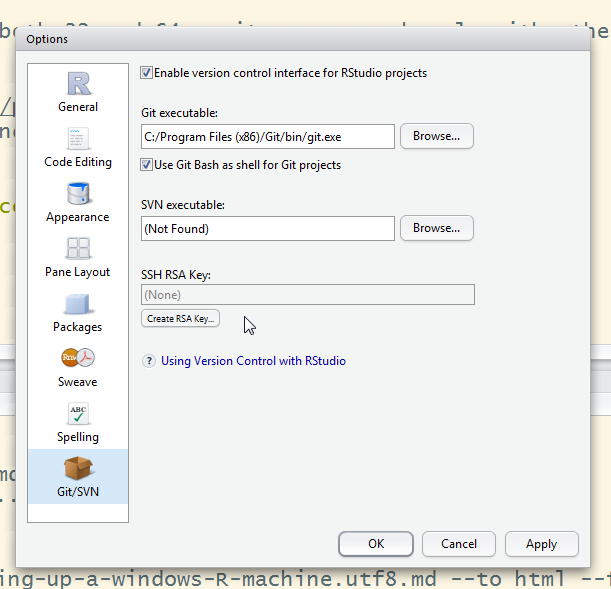
git config --global user.name="Your name"git config --global user.email="email@addre.ss"r
# Case-sensitive!
devtools::install_github("stephlocke/Rtraining")
https://github.com/stephlocke/Rtraining.git| Action | Operator | Example |
|---|---|---|
| Subtract | - | 5 - 4 = 1 |
| Add | + | 5 + 4 = 9 |
| Multiply | * | 5 * 4 = 20 |
| Divide | / | 5 / 4 = 1.25 |
| Raise to the power | ^ | 5 ^ 4 = 625 |
| Modulus | %% | 10 %% 4 = 2 |
| Absolute remainder | %/% | 9 %/% 4 = 2 |
| Basic sequence | : | sum(1:3) = 6 |
| Action | Operator | Example |
|---|---|---|
| Less than | < | 5 < 5 = FALSE |
| Less than or equal to | <= | 5 <= 5 = TRUE |
| Greater than | > | 5 > 5 = FALSE |
| Greater than or equal to | >= | 5 >= 5 = TRUE |
| Exactly equal | == | (0.5 - 0.3) == (0.3 - 0.1) is FALSE, 2 == 2 is TRUE |
| Not equal | != | (0.5 - 0.3) != (0.3 - 0.1) is TRUE, 2 != 2 is FALSE |
| Equal | all.equal() | all.equal(0.5 - 0.3,0.3 - 0.1) is TRUE |
| States | Representation |
|---|---|
| True | TRUE 1 |
| False | FALSE 0 |
| Empty | NULL |
| Unknown | NA |
Not a number e.g. 0/0 |
NaN |
Infinite e.g. 1/0 |
Inf |
| Action | Operator | Example |
|---|---|---|
| Not | ! | !TRUE is FALSE |
| And | & | TRUE & FALSE is FALSE, c(TRUE,TRUE) & c(FALSE,TRUE) is FALSE, TRUE |
| Or | ||
| Xor | xor() | xor(TRUE,FALSE) is TRUE |
| Bitwise And | && | c(TRUE,TRUE) && c(FALSE,TRUE) is FALSE |
| Bitwise Or | ||
| In | %in% | "Red" %in% c("Blue","Red") is TRUE |
| Not in | !( x %in% y) or Hmisc::%nin% | "Red" %nin% c("Blue","Red") = FALSE |
| Type | Implementation | Example |
|---|---|---|
| If | if(condition) {dosomething} | if(TRUE) { 2 } is 2 |
| If else | if(condition) {do something} else {do something different} or ifelse(condition, do something, do something else) | if(FALSE) { 2 } else { 3 } is 3 ifelse(FALSE, 2, 3) is 3 |
| For loop | for(i in seq) {dosomething} or foreach::foreach(i=1:3) %do% {something} | foreach(i=1:3) %do% {TRUE} is TRUE, TRUE, TRUE |
| While loop | while(condition) {do something } | a<-0 ; while(a<3){a<-a+1} ; a is 3 |
| Switch | switch(value, …) | switch(2, "a", "b") is b |
| Case | memisc::cases(…) | cases("pi<3"=pi<3, "pi=3"=pi==3,"pi>3"=pi>3) is pi>3 |
NB: If you find yourself using a loop, there's probably a better, faster solution
| Action | Operator | Example |
|---|---|---|
| Create / update a variable | <- | a <- 10 |
NB: There are others you could use, but this is the best practice
| Action | Operator | Example |
|---|---|---|
| Use public function from package | :: | memisc::cases() |
| Use private function from package | ::: | optiRum:::pounds_format() |
| Get a component e.g a data.frame column | $ | iris$Sepal.Length |
| Extract a property from a class | @ |
Won't be used in this course |
| Refer to positions in a data.frame or vector | [ ] | iris[5:10,1] |
| Refer to item in a list | [[ ]] | list(iris=iris,mtcars=mtcars)[["iris"]] |
| Action | Operator | Example |
|---|---|---|
| Comment | # | # This is my comment |
| Help | ? | ?data.table |
| Identifier | irisDT[ , `:=`(CreatedDate = Sys.Date())] |
| Data type | Example |
|---|---|
| Integer | 1 |
| Logical | TRUE |
| Numeric | 1.1 |
| String / character | “Red” |
| Factor (enumerated string) | “Amber” or 2 in c(“Red”,“Amber”,“Green”) |
| Complex | i |
| Date | “2015-04-24” |
| Data type | Info | Construction example(s) |
|---|---|---|
| Vector | A 1D set of values of the same data type | c(1,“a”) , 1:3 , LETTERS |
| Matrix | A 2D set of values of the same data type | matrix(LETTERS,nrow=13, ncol=2) , rbind(1:5,2:6) |
| Array | An nD set of values of the same data type | array(LETTERS, c(13,2)) |
| Data.frame | A 2D set of values of different data types | data.frame(a=1:26, b=LETTERS) |
| List | A collection of objects of various data types | list(vector=c(1,“a”), df=data.frame(a=1:6)) |
| Classes | A class is like a formalised list and can also contain functions i.e. methods | Won't be covered in this class |
NB: Most of my work uses vectors, data.tables (a souped up version of data.frames), and lists
| Function | Use |
|---|---|
| is.[data type] | Whether a vector is of a particular type |
| as.[data type] | Attempts to coerce a vector to a data type |
| str | Structure of an object including class/data type, dimensions |
| class | The class(es)/data type(s) an object belongs to |
| summary | Summarises an object |
| dput | Get R code that recreates an object |
| unlist | Simplify a list to a vector |
| dim | Dimensions of a data type |
| Format | Functions |
|---|---|
| CSV | read.csv , data.table::fread , readr::read_csv |
| Excel | readxl::read_excel |
| Database | RODBC::sqlQuery , DBI::dbGetQuery |
| SPSS / SAS / Stata | haven::read_[prog] |
| Format | Functions |
|---|---|
| CSV | write.csv |
| Excel | openxlsx::write.xlsx |
| Database | RODBC::sqlSave , DBI::dbWriteTable |
| SPSS / SAS / Stata | foreign::write.foreign |
As well standard formats, there's a lot of connector packages out there, including a suite for Hadoop.
DT[i, j, by]
DT[WHERE | JOIN | ORDER, SELECT | UPDATE, GROUP]
A data.table acts like an in-memory RDBMS:
There are some differences that need to be mentioned:
| Task | Generic syntax | Example(s)* |
|---|---|---|
| CREATE | data.table(…) setDT() | data.table(a=1:3 , b=LETTERS[1:3]) data.table(iris) |
| PRIMARY KEY | data.table(…,key) setkey() | data.table(a=1:3 , b=LETTERS[1:3], key="b") setkey(data.table(iris),Species) |
| SELECT basic | DT[ , .( cols )] | irisDT[ , .(Species, Sepal.Length)] |
| SELECT alias | DT[ , .( a=col )] | irisDT[ , .(Species, Length=Sepal.Length)] |
| SELECT COUNT | DT[ , .N] | irisDT[ ,.N] |
| SELECT COUNT DISTINCT | DT[ , uniqueN(cols)] | irisDT[ ,uniqueN(.SD)] |
| SELECT aggregation | DT[ , .( sum(col) , .N )] | irisDT[ , .(Count=.N, Length=mean(Sepal.Length))] |
| WHERE exact on primary key | DT[value] DT[value, ] | irisDT["setosa"] irisDT["setosa", .(Count=.N)] |
| WHERE | DT[condition] DT[condition, j, by] | irisDT[Species=="setosa"] irisDT[Species=="setosa", .(Count=.N)] |
| WHERE BETWEEN | DT[between(col, min, max)] DT[ col %between% c(min,max) ] | irisDT[between(Sepal.Length, 1, 5)] irisDT[Sepal.Length %between% c(1,5)] |
| WHERE LIKE | DT[like(col,pattern)] DT[ col %like% pattern ] | irisDT[like(Species,"set")] irisDT[Species %like% "set"] |
| ORDER asc. | DT[order(cols)] DT[order(cols), j, by] | irisDT[order(Species)] |
| ORDER desc. | DT[order(-cols)] DT[order(-cols), j, by] | irisDT[order(-Species)] |
| ORDER multiple | DT[order(cols)] DT[order(cols), j, by] | irisDT[order(-Species, Petal.Width)] |
| GROUP BY single | DT[i, j, by] | irisDT[ ,.N, by=Species] |
| GROUP BY multiple | DT[i, j, by] | irisDT[ ,.N, by=.(Species,Width=Petal.Width)] |
| TOP | head(DT, n) | head(irisDT) |
| HAVING | DT[i, j, by][condition] | irisDT[ , .(Count=.N), by=Species][Count>25] |
| Sub-queries | DT[…][…][…] | irisDT[ , .(Sepal.Length=mean(Sepal.Length)), by=Species][Sepal.Length>6, .(Species)] |
* Uses irisDT <- data.table(iris)
| Task | Generic syntax | Example(s)* |
|---|---|---|
| INSERT | DT <- rbindlist(DT, newDT) | irisDT<-rbindlist( irisDT, irisDT[1] ) |
| READ aka SELECT (see above) | DT[ , .( cols )] | irisDT[ , .(Species, Sepal.Length)] |
| UPDATE / ADD column | DT[ , a := b ] | irisDT[ , Sepal.Area := Sepal.Width * Sepal.Length] |
| UPDATE / ADD multiple columns | DT[ , :=(a = b, c = d) ] | irisDT[ , `:=`(CreatedDate = Sys.Date(), User = "Steph")] |
| UPDATE / ADD multiple columns by reference | DT[ , (newcols):=vals ] | irisDT[ , c("a","b"):=.(1,2)] |
| DELETE | DT <- DT[!condition] | irisDT <- irisDT[!(Species=="setosa" & Petal.Length>=1.5)] |
| DROP table | DT <- NULL | irisDT<-NULL |
| DROP column | DT[,col:=NULL] | iristDT[,Species:=NULL] |
* Uses irisDT <- data.table(iris)
| Task | Generic syntax | Example(s)* |
|---|---|---|
| Structure | str(DT) | str(irisDT) |
| Column Names | colnames(DT) | colnames(irisDT) |
| Summary stats | summary(DT) | summary(irisDT) |
| Retrieve primary key info | key(DT) | key(irisDT) |
| List all data.tables | tables() | tables() |
* Uses irisDT <- data.table(iris)
| Task | Generic syntax | Example(s)* |
|---|---|---|
| INNER JOIN | Y[X, nomatch=0] | lookupDT[irisDT,nomatch=0] |
| LEFT JOIN | Y[X] | lookupDT[irisDT] |
| FULL JOIN | merge(X, Y, all=TRUE) | merge(irisDT, lookupDT, all=TRUE) |
| CROSS JOIN | optiRum::CJ.dt(X,Y) | CJ.dt(irisDT, lookupDT) |
| UNION ALL | rbindlist( list(X,Y), fill=TRUE ) | rbindlist( list(irisDT, lookupDT), fill=TRUE ) |
| UNION | unique( rbindlist( list(X,Y), fill=TRUE ) ) | unique( rbindlist( list(irisDT, lookupDT), fill=TRUE ) ) |
| JOIN and AGGREGATE | Y[X, cols, by] | lookupDT[irisDT,.(count=.N),by=Band] |
* Uses:
irisDT <- data.table(iris, key="Species")
lookupDT <- data.table(Species=c("setosa", "virginica", "Blah"), Band=c("A", "B", "A"), key="Species")
| Task | Generic syntax | Example(s)* |
|---|---|---|
| SELECT dynamically | DT[ , colnames , with=FALSE] , DT[ , .SD , .SDcols=colnames | cols<-colnames(irisDT); irisDT[ , cols, with=FALSE] cols<-colnames(irisDT); irisDT[ , .SD, .SDcols=colnames] |
| GROUP BY dynamically | DT[ , …, by=colnames] | irisDT[,.N,by=c("Species")] |
| GROUP BY, ORDER BY group | DT[ , … , keyby] | irisDT[,.N,keyby=Species] |
| UPDATE / ADD column of summary stat | DT[ , a := b ] | irisDT[ , All.SL.Mean:=mean(Sepal.Length)] |
| UPDATE / ADD column by group | DT[ , a := b, by] | irisDT[ , Species.SL.Mean:=mean(Sepal.Length), by=Species] |
| TOP by group | DT[ , head(.SD), by] | irisDT[ , head(.SD,2) , by=Species] |
| Largest record | DT[ which.max(col) ] | irisDT[ which.max(Sepal.Length) ] |
| Largest record by group | DT[ , .SD[ which.max(col) ], by] | irisDT[ , .SD[ which.max(Sepal.Length) ], by=Species] |
| Cumulative total | DT[ , cumsum(col) ] | irisDT[ , cumsum(Sepal.Width)] |
| NEGATIVE SELECT | DT[ , .SD, .SDcols=-“colname”] | irisDT[ , .SD, .SDcols=-"Species"] |
| RANK | DT[ , frank(col) ] | irisDT[ , frank(Sepal.Length,ties.method="first")] |
| AGGREGATE multiple columns | DT[ , lapply(.SD, sum)] | irisDT[ , lapply(.SD,sum), .SDcols=-"Species"] |
| AGGREGATE multiple columns by group | DT[ , lapply(.SD, sum), by] | irisDT[ , lapply(.SD,sum), by=Species] |
| COUNT DISTINCT multiple columns by group | DT[ , lapply(.SD, uniqueN), by] | irisDT[ , lapply(.SD,uniqueN), by=Species] |
| COUNT NULL multiple columns by group | DT[ , lapply(.SD, function(x) sum(is.na(x))), by] | irisDT[ , lapply(.SD,function(x) sum(is.na(x))), by=Species] |
| PIVOT data - to single value column | melt(DT,…) | melt(irisDT) |
| PIVOT data - to aggregate | dcast(DT, a~b, function) | dcast(melt(irisDT), Species ~ variable, sum) |
| Convert a large data.frame or list | setDT() | iris<-iris; setDT(iris) |
| ROW_NUMBER | DT[ , .I] | irisDT[ , .I] |
| GROUP number | DT[, .GRP ,by] | irisDT[ , .GRP, by=Species] |
* Uses irisDT <- data.table(iris)
| Task | Generic syntax | Example(s)* |
|---|---|---|
| GROUP BY each new incidence of group | DT[ , cols , by=(col, rleid(col))] | irisDT[order(Sepal.Length), .N, by=.(Species, rleid(Species))] |
| Calculate using (previous/next) N row | DT[ , shift( cols, n)] | irisDT[ , prev.Sepal.Length:=shift(Sepal.Length), by=Species ] |
| ORDER underlying table | setorder() | setorder(irisDT,Species) |
| JOIN & GROUP by keys | X[Y, .N, by=.EACHI] | irisDT[lookupDT, .N, by=.EACHI] |
| TRANSPOSE data.table | transpose(DT) | transpose(irisDT) |
| Split string to columns | DT[, tstrsplit(charCol, pattern) ] | irisDT[ , tstrsplit(as.character(Species),"e")] |
* Uses:
irisDT <- data.table(iris, key="Species")
lookupDT <- data.table(Species=c("setosa", "virginica", "Blah"), Band=c("A", "B", "A"), key="Species")
This intro covers the charting package ggplot2.
The “base” charting functionality will not be covered because it's much more difficult to achieve good looking results quickly and I don't believe in that much effort for so little benefit!
ggplot2 is a plotting system for R, based on the grammar of graphics, which tries to take the good parts of base and lattice graphics and none of the bad parts. It takes care of many of the fiddly details that make plotting a hassle (like drawing legends) as well as providing a powerful model of graphics that makes it easy to produce complex multi-layered graphics.
| Term | Explanation | Example(s) |
|---|---|---|
| plot | A plot using the grammar of graphics | ggplot() |
| aesthetics | attributes of the chart | colour, x, y |
| mapping | relating a column in your data to an aesthetic | |
| statistical transformation | a translation of the raw data into a refined summary | stat_density() |
| geometry | the display of aesthetics | geom_line(), geom_bar() |
| scale | the range of values | axes, legends |
| coordinate system | how geometries get laid out | coord_flip() |
| facet | a means of subsetting the chart | facet_grid() |
| theme | display properties | theme_minimal() |
library(ggplot2)
p <- ggplot(data=iris)
p <- ggplot(data=iris, aes(x=Sepal.Width, y=Sepal.Length, colour=Species))
p <- p + geom_point()
p
p <- p + stat_boxplot(fill="transparent")
p
## Warning: position_dodge requires non-overlapping x intervals
p <- p + coord_flip()
p
## Warning: position_dodge requires non-overlapping x intervals
p <- p + facet_grid(.~Species)
p
p <- p + optiRum::theme_optimum()
p
ggplot(data=iris, aes(x=Sepal.Width, y=Sepal.Length, colour=Species)) +
geom_point() +
stat_boxplot(fill="transparent") +
# coord_flip() + # Commented out
facet_grid(.~Species) +
optiRum::theme_optimum()
Producing documents / documentation directly in R means that you closely interweave (knit) your analysis and R code together. This reduces rework time when you want to change or extend your code, it reduces time to produce new versions, and because it's code it's easier to apply strong software development principles to it.
Oh, and you don't need to spend hours making text boxes in powerpoint! Win ;-)
There are two languages which you can knit your r code into:
Markdown is great for very quick generation and light (or css driven) styling and is what this section focusses on. LaTeX is excellent for producing stunning, more flexible documents.
The following text is the default text that gets created when you produce a new rmarkdown file in rstudio
This is an R Markdown document. Markdown is a simple formatting syntax for authoring HTML, PDF, and MS Word documents. For more details on using R Markdown see http://rmarkdown.rstudio.com.
When you click the Knit button a document will be generated that includes both content as well as the output of any embedded R code chunks within the document. You can embed an R code chunk like this:
summary(cars)
## speed dist
## Min. : 4.0 Min. : 2.00
## 1st Qu.:12.0 1st Qu.: 26.00
## Median :15.0 Median : 36.00
## Mean :15.4 Mean : 42.98
## 3rd Qu.:19.0 3rd Qu.: 56.00
## Max. :25.0 Max. :120.00
You can also embed plots, for example:
Note that the echo = FALSE parameter was added to the code chunk to prevent printing of the R code that generated the plot.
The following text is part of the standard documentation on rmarkdown. I pull it from github.com/rstudio/rmarkdown and integrate it using knitr. It is better than I could produce and the act of integrating it gives an extra example of useful ways to build documents.
This document provides quick references to the most commonly used R Markdown syntax. See the following articles for more in-depth treatment of all the capabilities of R Markdown:
*italic* **bold**
_italic_ __bold__
# Header 1
## Header 2
### Header 3
Unordered List:
* Item 1
* Item 2
+ Item 2a
+ Item 2b
Ordered List:
1. Item 1
2. Item 2
3. Item 3
+ Item 3a
+ Item 3b
R code will be evaluated and printed
```{r}
summary(cars$dist)
summary(cars$speed)
```
There were 50 cars studied
Use a plain http address or add a link to a phrase:
http://example.com
[linked phrase](http://example.com)
Images on the web or local files in the same directory:


A friend once said:
> It's always better to give
> than to receive.
Plain code blocks are displayed in a fixed-width font but not evaulated
```
This text is displayed verbatim / preformatted
```
We defined the `add` function to
compute the sum of two numbers.
LaTeX Equations
Inline equation:
$equation$
Display equation:
$$ equation $$
Three or more asterisks or dashes:
******
------
First Header | Second Header
------------- | -------------
Content Cell | Content Cell
Content Cell | Content Cell
Reference Style Links and Images
A [linked phrase][id].
At the bottom of the document:
Images
![alt text][id]
At the bottom of the document:
End a line with two or more spaces:
Roses are red,
Violets are blue.
superscript^2^
~~strikethrough~~
library(data.table)
library(shiny)
defaultdisplay<-list(
width="100%", height="75%"
)
shinyAppDir(
system.file("examples/06_tabsets", package="shiny"),
options = defaultdisplay
)
A shiny application report consists of two functions:
shinyServer()shinyUI()One says what to execute and the other states how to present it. Do all data manipulation, chart production in shinyServer()
defaultdisplay<-list(width="100%", height="75%")
shinyApp(
ui = fluidPage(),
, server = function(input, output) {}
, options = defaultdisplay
)
You typically split into two files:
shinyServer()shinyUI()This can then be run with runApp()
You can do a single file example app.R which contains both functions but this is typically better for very short apps.
Use these just inside shinyUI() to produce a layout
## Page Types
## 1: basicPage
## 2: bootstrapPage
## 3: fillPage
## 4: fixedPage
## 5: fluidPage
## 6: navbarPage
## 7: updateNavbarPage
shinyApp(
ui = fluidPage(dateInput("datePicker", "Pick a date:",
format="dd/mm/yy"),
dateRangeInput("dateRange", "Pick dates:",
start=Sys.Date(),
end=Sys.Date() ) ),
server = function(input, output) {}
,options = defaultdisplay
)
Basic
shinyApp(
ui = fluidPage(numericInput("vals", "Insert a number:",
value=15, min=10) ),
server = function(input, output) {}
,options = defaultdisplay
)
Sliders
shinyApp(
ui = fluidPage(sliderInput("vals", "Insert a number:",
min=0, max=50, value=15) ),
server = function(input, output) {}
,options = defaultdisplay
)
A single line
shinyApp(
ui = fluidPage(textInput("char", "Insert text:") ),
server = function(input, output) {}
,options = defaultdisplay
)
A paragraph
shinyApp(
ui = fluidPage(tags$textarea(id="charbox", rows=3,
cols=40, "Default value") ),
server = function(input, output) {}
,options = defaultdisplay
)
shinyApp(
ui = fluidPage(selectInput("multiselect", "Pick favourites:",
c("Green","Red","Blue"),
multiple=TRUE) ),
server = function(input, output) {}
,options = defaultdisplay
)
## Input controls
## 1: checkboxGroupInput
## 2: checkboxInput
## 3: dateInput
## 4: dateRangeInput
## 5: fileInput
## 6: numericInput
## 7: passwordInput
## 8: registerInputHandler
## 9: removeInputHandler
## 10: restoreInput
## 11: selectInput
## 12: selectizeInput
## 13: sliderInput
## 14: textAreaInput
## 15: textInput
## 16: updateCheckboxGroupInput
## 17: updateCheckboxInput
## 18: updateDateInput
## 19: updateDateRangeInput
## 20: updateNumericInput
## 21: updateSelectInput
## 22: updateSelectizeInput
## 23: updateSliderInput
## 24: updateTextAreaInput
## 25: updateTextInput
## Input controls
shinyApp(
ui = fluidPage(textInput("char", "Insert text:") ,
textOutput("text") ),
server = function(input, output) {
output$text <- renderText(input$char)
} ,options = defaultdisplay
)
shinyApp(
ui = fluidPage(tableOutput("basictable") ),
server = function(input, output) {
output$basictable <- renderTable(head(iris,5))
} ,options = defaultdisplay
)
shinyApp(
ui = fluidPage(dataTableOutput("datatable") ),
server = function(input, output) {
output$datatable <- renderDataTable(head(iris,5))
} ,options = defaultdisplay
)
shinyApp(
ui = fluidPage(plotOutput("chart") ),
server = function(input, output) {
output$chart <- renderPlot(pairs(iris))
} ,options = defaultdisplay
)
a <- reactive({input$a})
a
shinyApp(
ui = fluidPage(textInput("char", "Insert text:") ,
textOutput("textA"),textOutput("textB") ),
server = function(input, output) {
char<-reactive({rep(input$char,5)})
output$textA <- renderText(paste(char(),collapse="+"))
output$textB <- renderText(paste(char(),collapse="-"))
}
,options = defaultdisplay
)
shinythemesrvestshiny::runApp()shinyApps packagePronunciation: lay-tech
LaTeX is an open source markup language with a typesetting engine. It's been around since the 70s and generally makes awesome documents.
LaTeX is designed to work stand-alone, or integrate with other languages. It's particularly good with R.
It works in the way rmarkdown does with knitr, but allows for more sophisticated document styling.
sudo apt-get texliveOnce you have a LaTeX installation, you can write LaTeX in Rstudio.
In Rstudio, select the File type “R Sweave” which saves as a .Rnw file.
LaTeX is a very deep language, so no attempt is made to teach you here.
One of the easiest ways of getting started with LaTeX (generally) is to pick one of the examples / templates on overleaf.com and play with it.
For using R and LaTeX, you can use the minimal examples on yihui.name/knitr to get started.
See the LaTeX wikibook for lots of info about LaTeX.

SOURCE CONTROL IS FOR ALL THE THINGS
Source control is important because it provides:
There are two types of source control systems:
Centralised means that there is a single storage location and to work on a file it must be exclusively checked out. Distributed systems involve taking copies of the code base, making changes, and pushing these back to primary storage location.
Both have their own disadvantages but since with distributed source control you never get the situation where someone's left a file checked out as they go on holiday and no-one else can use it, I'm a big fan of distributed source control systems.
Git is a distributed source control system.
It integrates neatly into Rstudio, making it easy to source control your analysis.
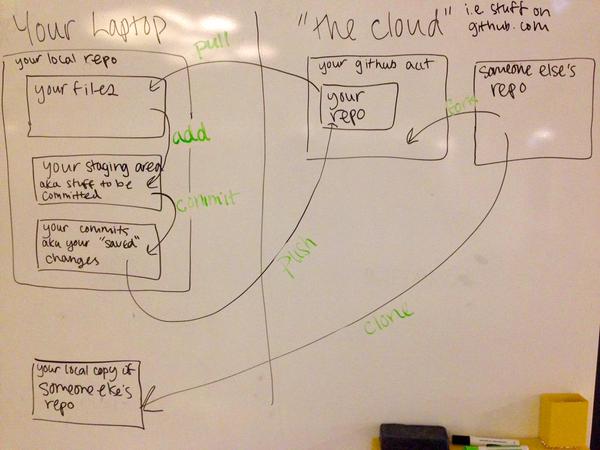
There are more terms. For a friendly glossary see Github's git glossary, and for an extensive, technical glossary see the official Git glossary
The package git2r supports a source control workflow directly within R. This means you can continue to use Rstudio for even complex git tasks. And of course there's always the shell option in Rstudio.
For a handy Git cheatsheet, check out this GitHub one.
The git2r documentation is pretty good. It's easier though to use once you've been utilising the Rstudio GUI for a bit, and dabbling with the command line.
A package is a collection of functionality designed to achieve one or more purposes. Commonly it is a bundle of functions that help tackle a certain type of analysis.
Packages are great ways to modularise your code and create standardised ways of doing specific tasks in your organisation, like charts (optiRum::theme_optimum()).
There is an R foundation guide to writing packages. I don't recommend you start with that! It is however what any package that you submit to the central repository of R packages (CRAN) will be held against - so if you'd like to get a package on CRAN you will need to read this.
The better, more accessible book R packages is by Hadley Wickham and will cover things in a lot of depth but is more accesable and has exercises.
For quick learning abotu devtools you can check out the cheatsheet
The easiest way to build a good quality package is to use the package devtools. This is a package designed specifically to make life easier for package developers.
Here is my typical workflow:
library(devtools)
pkg<-"newPackage"
create(pkg)
# Open the project!
library(devtools)
# Add unit test framework
add_test_infrastructure()
# Add CI framework
add_travis()
# Add folder for macro-level help files
use_vignette()
# Add file for providing info about your package
use_package_doc()
# Add a file for storing comments about the release if submitting to CRAN
use_cran_comments()
# Create various useful files
file.create("README.md")
file.create("NEWS")
# Set git up
library(git2r)
init(".")
Once I have this skeleton I fill in the various bits of info about my package in DESCRIPTION, README, R/package.R, and so forth.
After I've done some basic hygiene, I can start building my R functions and associated tests.
...) argument to pass values through to optional components in functions#' A function quick description
#'
#' A more detailed description that can span multiple lines
#' for readability. Covers concepts, typical usages etc.
#'
#' @param param1 Info about param1 e.g. data type, guidance
#' @param param2 Info about param1 e.g. data type, guidance
#' @param ... Additional values to pass to x, y, z
#'
#' @return returnDT Info about what is returned by the function
#'
#' @keywords words allowing search
#' @family ifPartOfABundle
#'
#' @examples
#' # Sample code that illustrates usage
#'
#' @export
myFuncName<- function(param1, param2="Blah", ...){
stopifnot(param1>0, is.character(param2))
# Function code routinely commented with WHY or
# explanations of complex HOW (but consider
# breaking these up / simplifying)
}
Let's make sure your code is all working (assumes you've got unit tests)
library(devtools)
# Build help files
document()
# Run unit tests
test()
# Check against CRAN standards
check()
Each of these steps could identify things to fix. It's great to get rid of as many ERRORs, WARNINGs, and NOTEs as possible.
There are a number of locations but I'll cover two:
My personal recommendations are to use GitHub as your active development environment, so that people can download the latest version, and periodically attempt to release to CRAN. This helps push up your package quality and makes your code more widely available.
Unit testing is the technique of writing tests that assess low-level functionality against requirements.
In reports, databases, and code we typically encode business rules, conventions, and our own conventions. Bringing these into reusable functions means less code reproduction, no variances between team members, and lower time to change.
All of these are verifiably correct and so can be tested. Therefore, why test them manually, or why risk someone “tweaking” the code and changing the rules without people knowing? Your unit tests save you time over the long run, and protect against unexpected behaviour changes.
Reconciliations can also be done inside your unit tests. If you have “canonical answers”, you can test new transformations against these to ensure you're consistent.
I'm not a developer so this may be the wrong approach but here are the scenarios I write tests for:
Also, if you're struggling to test because it's got really complex inputs, outputs, or intermediate calculations, consider breaking up the code and rewriting. It's more likely that things are going to go wrong and you won't know if something is particularly complex.
Let's build a sample function:
myfunc<-function(a=1,b=2,c="blah"){
stopifnot(is.numeric(a), is.numeric(b),
is.character(c))
d<-if(a<0){
a*b
}else{
a/b
}
e<-paste0(c,d)
return(e)
}
Let's write some tests (in a file tests/testthat/test-myfunc.r)
# Add a high-level name for group of tests, typically the function name
context("myfunc")
# Simplest test
test_that("Defaults return expected result",{
result<-myfunc()
check<-"blah0.5"
expect_equal(result,check)
})
# Vector test
test_that("Basic vectorisation works",{
result<-myfunc(a=c(1,1),b=c(2,2), c=c("blah","blah"))
check<-c("blah0.5","blah0.5")
expect_equal(result,check)
})
# Non-uniform vectorisation test
test_that("Complex vectorisation works",{
result<-myfunc(a=c(1,1),b=c(2,2))
check<-c("blah0.5","blah0.5")
expect_equal(result,check)
})
# Test a different condition
test_that("Negative a values result in multiplication",{
result<-myfunc(a=c(-1,-2),b=c(2,2))
check<-c("blah-2","blah-4")
expect_equal(result,check)
})
# Test a different condition
test_that("a=0 values result in 0",{
result<-myfunc(a=0)
check<-c("blah0")
expect_equal(result,check)
})
# Test some duff inputs
test_that("errors expectedly",{
expect_error(myfunc(a="0"))
expect_error(myfunc(b="0"))
expect_error(myfunc(c=0))
})
There are a lot more expectation functions you can use and you can make your own.
With excellent guidance and tooling on making R packages, it's becoming really easy to make a package to hold your R functionality. This has a host of benefits, not least source control (via GitHub) and unit testing (via the testthat package). Once you have a package and unit tests, a great way of making sure that as you change things you don't break them is to perform Continuous integration.
What this means is that every time you make a change, your package is built and thoroughly checked for any issues. If issues are found the “build's broke” and you have to fix it ASAP.
The easiest, cheapest, and fastest way of setting up continuous integration for R stuff is to use Travis-CI, which is free if you use GitHub as a remote server for your code. NB - it doesn't have to be your only remote server
The first thing that needs doing is setting up your accounts and turning on CI for your repositories. The website is pretty good so I won't go into a lot of detail, but the process is:
Additionally, whilst we're doing this we should be awesome and set up test coverage checks as well. The process is really similar, but for coveralls.io and we only need the one set of config details in our package.
Then you add a really simple file into the root of your project called .travis.yml.
This should contain, at minimum, the following:
language: r
sudo: required
r_github_packages:
- jimhester/covr
after_success:
- Rscript -e 'library(covr);coveralls()'
NB - be careful with the indentation, YAML is very sensitive!
This is the latest set of values that work as it takes into account the recent support for R, the ability to reference github packages, and also Travis' move towards docker containers which don't accept sudo commands.
Once you've flipped the switch on Travis and Coveralls, every push to GitHub will trigger Travis. Travis will basically build a server with all the requirements needed to run R and build R packages. It'll then install all your package's dependencies, check the package for minimum quality standards and also run your testthat tests. Once this is done the final bit tests your code coverage and passes the results to Coverall.
Great, so you've checked the sites and it's working but you should show the world it's working! You can get some some snippets of code from each of the sites that you can paste into your README file. These stay up to date with the latest results so that you (and everyone else) can see the status of your package.
Azure portal, using gallery creation for VM
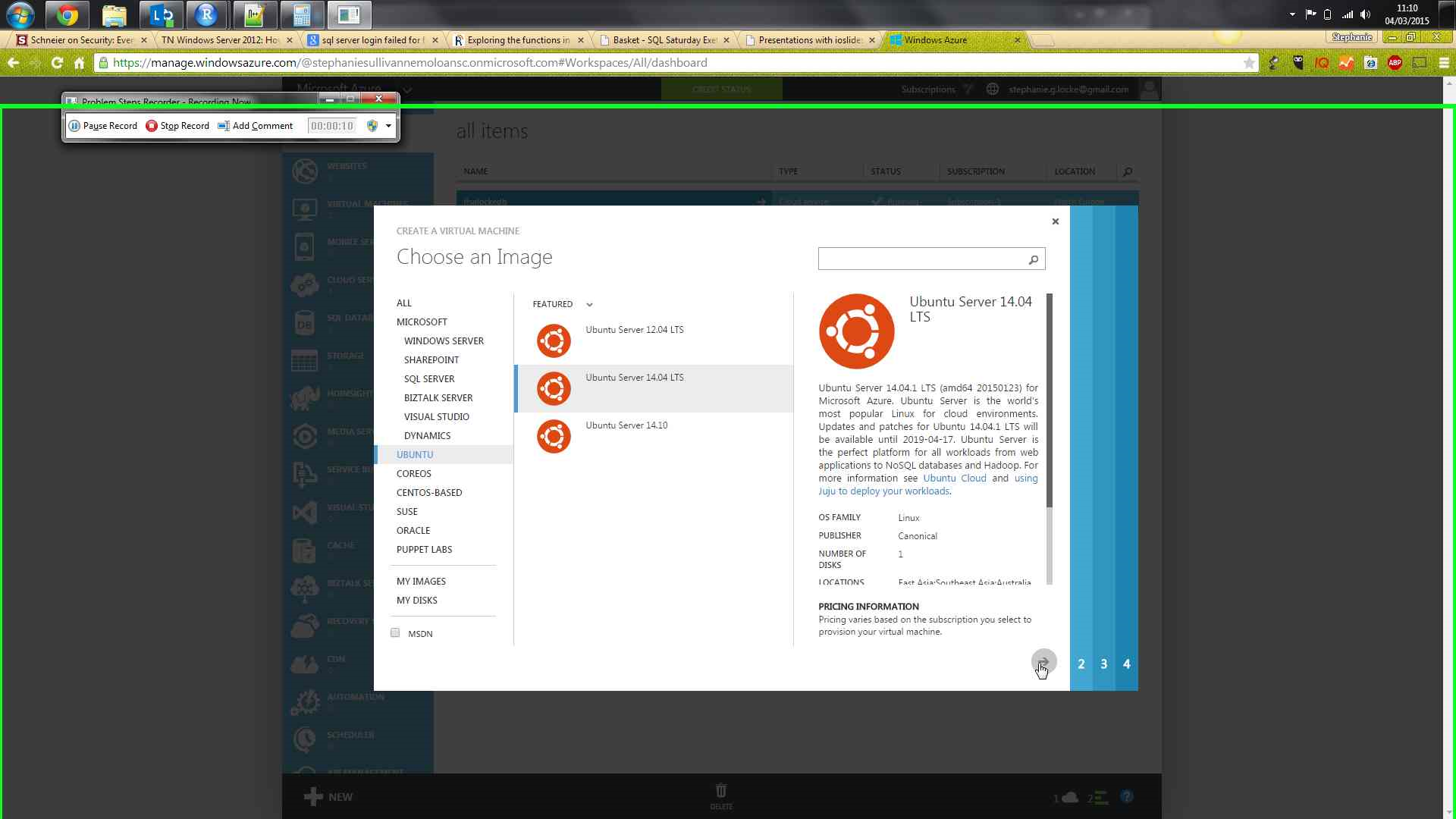
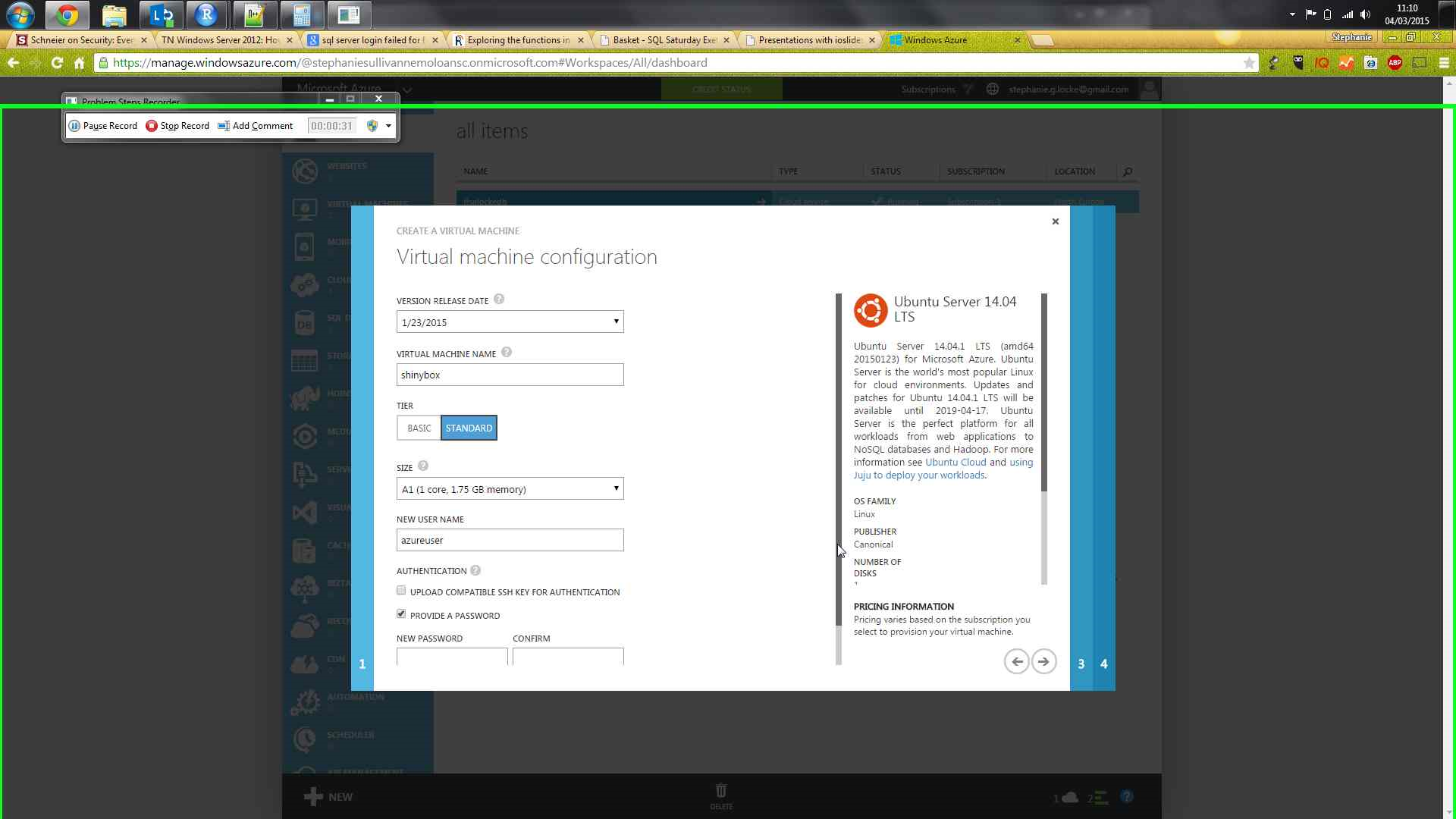
sudo apt-get update to get the package repository metadatasudo apt-get install r-base to get R. Will have lots of extra associated packages - select Y when promptedsudo apt-get install gdebi-core to enable processing of rstudio installation packagesudo apt-get install libapparmor1 if using ubuntuwget http://download2.rstudio.org/rstudio-server-0.98.1103-amd64.debsudo gdebi rstudio-server-0.98.1103-amd64.deb
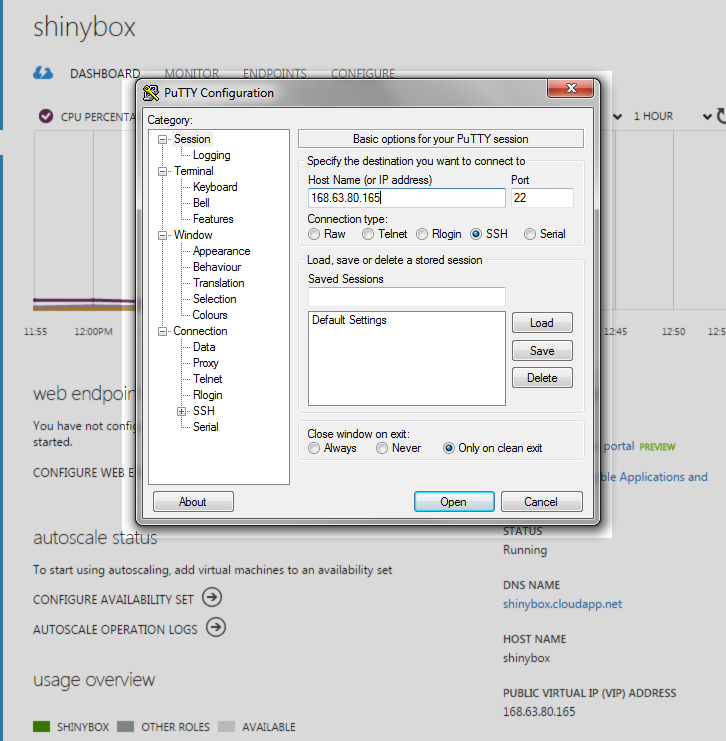
## Configuring port (away from 8787) and allowing on Azuresudo nano /etc/rstudio/rserver.confsudo rstudio-server restart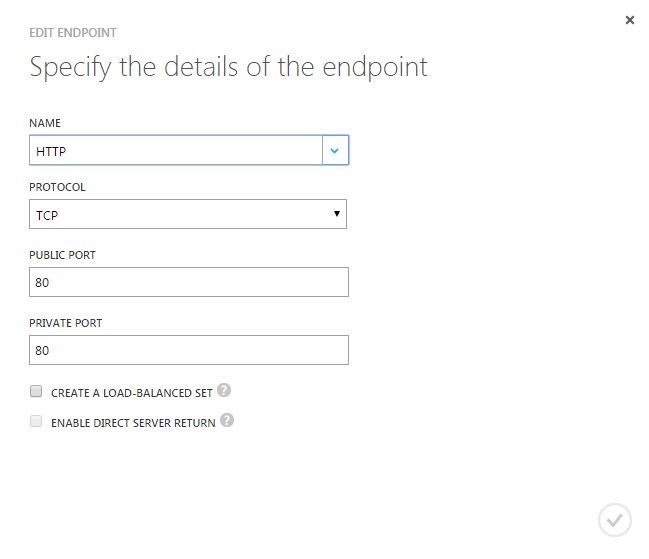
Azure portal, using gallery creation for VM



sudo apt-get update to get the package repository metadatasudo apt-get install r-base to get R. Will have lots of extra associated packages - select Y when promptedsudo su - -c "R -e \"install.packages('shiny', repos='http://cran.rstudio.com/')\"" to install shiny in Rsudo apt-get install gdebi-core to enable processing of shiny-server installation packagewget http://download3.rstudio.org/ubuntu-12.04/x86_64/shiny-server-1.3.0.403-amd64.debsudo gdebi shiny-server-1.3.0.403-amd64.debsudo nano /etc/shiny-server/shiny-server.confsudo restart shiny-server